Center for Wildlife Genetics
National Reference Center for Genetic Analysis of Wolf and Lynx
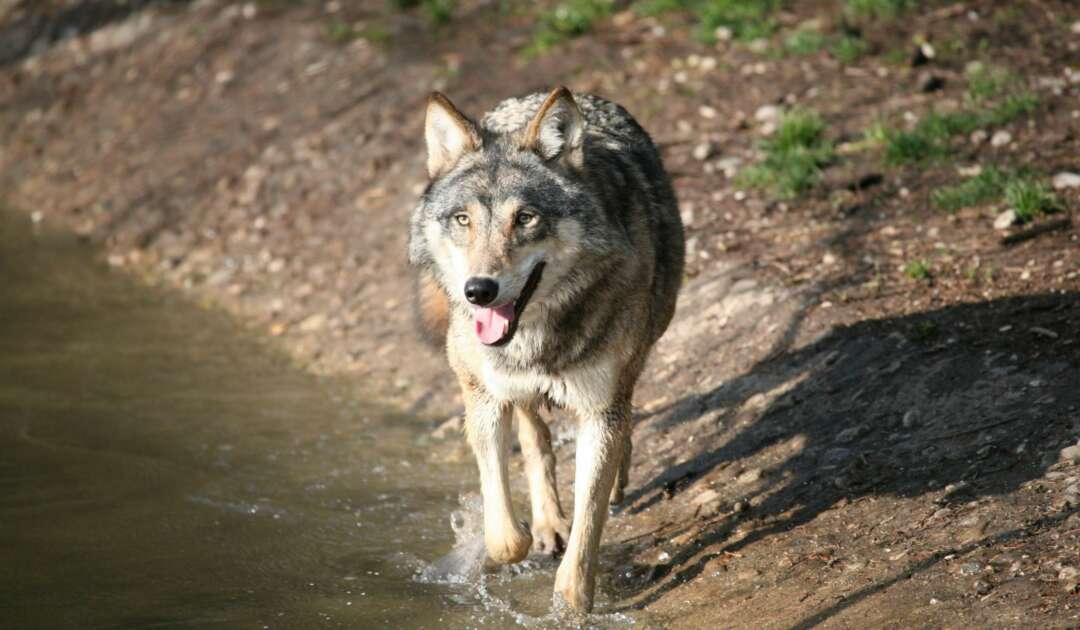
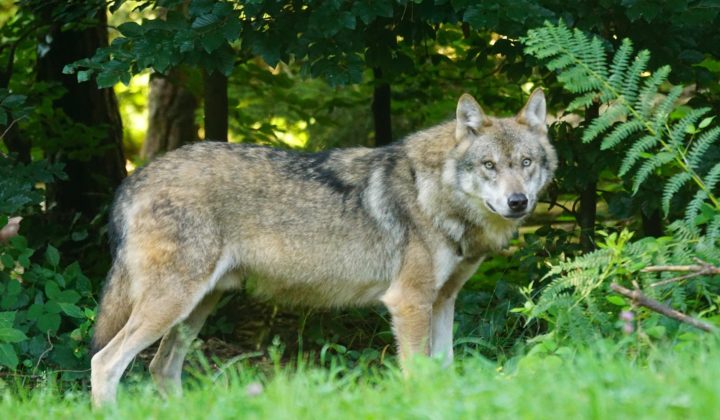
Since 2010, all samples collected in the frame of the German-wide wolf and lynx monitoring are centrally analyzed at the Senckenberg Center for Wildlife Genetics. The results are used to gain knowledge regarding the occurrence of wolves and lynx in Germany, to distinguish individuals, to determine relationships, and to continuously check the wolf population for possible admixture with domestic dogs (hybridization).
The DNA-based investigations of livestock depredations to detect wolves, lynx and other predators play an important role for monitoring and management. These are carried out on behalf of the German federal states, who are responsible for large carnivore monitoring. Accordingly, all results obtained are reported to the federal states.
Senckenberg was recommended to the federal states as a national reference center for genetic studies of wolves and lynx after a selection process supervised by the Federal Agency for Nature Conservation (BfN). The Federal working group on nature conservation, landscape management and recreation (LANA) decided in October 2009 to follow through with the recommendation of the BfN. The use of a central analysis laboratory ensures the comparability of all data accruing nationwide. This type of official wildlife monitoring utilizing a central processing center has become a well-established process internationally as a response to the current lack of standards available for genetic wildlife monitoring (de Groot et al. 2016).